Microbial interactions of zoonotic pathogens
We study human bacterial pathogens and their bacteriophages with focus on Staphylococcus aureus. We aim to understand how they respond to and survive adverse conditions including exposures to antibiotics, biocides and phages and how resistance may be mitigated.
Antibiotic resistance
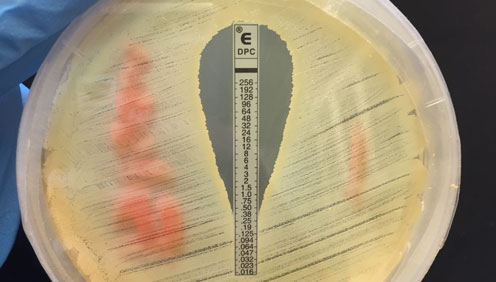
Antibiotic resistance can develop by accumulation of mutations as happens in S. aureus strains with decreased susceptibility to vancomycin, the VISA strains. We are interested in the process behind resistance development and how mutations in susceptible strains may prime bacteria for rapid resistance development.
Quorum sensing communication
Quorum sensing enables S. aureus to sense the density of S. aureus cells and control virulence gene expression accordingly. Biofilms may be a place where quorum sensing is particularly important. We search for compounds and organisms that interfere with S. aureus quorum sensing in order to develop new treatment options for serious staphylococcal infections.
Phages
We study the role of temperate phages in virulence of livestock strains of methicillin resistant S. aureus (MRSA) as well as quorum sensing control of phage infections. Also we investigate how antibiotic resistance impacts phage susceptibility as well as the transfer of resistance genes by phage transduction. See how in the movie Antibiotic Resistance Transfer in Staphylococcus aureus/Trends in Microbiology.
Yang J, Bowring JZ, Krusche J, Lehmann E, Bejder BS, Silva SF, Bojer MS, Grunert T, Peschel A, Ingmer H. Cross-species communication via agr controls phage susceptibility in Staphylococcus aureus. Cell Rep. 2023 Sep 26;42(9):113154. doi: 10.1016/j.celrep.2023.113154. Epub 2023 Sep 19. PMID: 37725513.
Fait A, Andersson DI, Ingmer H. Evolutionary history of Staphylococcus aureus influences antibiotic resistance evolution. Curr Biol. 2023 Aug 21;33(16):3389-3397.e5. doi: 10.1016/j.cub.2023.06.082. Epub 2023 Jul 25. PMID: 37494936.
Fait A, Seif Y, Mikkelsen K, Poudel S, Wells JM, Palsson BO, Ingmer H. Adaptive laboratory evolution and independent component analysis disentangle complex vancomycin adaptation trajectories. Proc Natl Acad Sci U S A. 2022 Jul 26;119(30):e2118262119. doi: 10.1073/pnas.2118262119. Epub 2022 Jul 19. PMID: 35858453; PMCID: PMC9335240.
Bowring JZ, Su Y, Alsaadi A, Svenningsen SL, Parkhill J, Ingmer H. Screening for Highly Transduced Genes in Staphylococcus aureus Revealed Both Lateral and Specialized Transduction. Microbiol Spectr. 2022 Feb 23;10(1):e0242321. doi: 10.1128/spectrum.02423-21. Epub 2022 Feb 9. PMID: 35138167; PMCID: PMC8826898.
Phage mediated bacterial transduction
Phage - antibiotic cooperativity
Quorum sensing control
Biocide susceptibility
Probiotic approaches to quorum sensing control
RNA single cell sequencing
Contact
Group Leader:
Prof Hanne Ingmer
Stigbøjlen 7
1870 Frederiksberg C
Ph: +45 35 33 27 73
Secretariat:
Nora Ottens
Grønnegårdsvej 15
Ph: +45 35 33 27 22





